Ben Goodrich writes:
The rstanarm R package, which has been mentioned several times on stan-users, is now available in binary form on CRAN mirrors (unless you are using an old version of R and / or an old version of OSX). It is an R package that comes with a few precompiled Stan models — which are called by R wrapper functions that have the same syntax as popular model-fitting functions in R such as glm() — and some supporting R functions for working with posterior predictive distributions. The files in its demo/ subdirectory, which can be called via the demo() function, show how you can fit essentially all of the models in Gelman and Hill’s textbook
http://stat.columbia.edu/~gelman/arm/
and rstanarm already offers more (although not strictly a superset of the) functionality in the arm R package.
The rstanarm package can be installed in the usual way with
install.packages(“rstanarm”)which does not technically require the computer to have a C++ compiler if you on Windows / Mac (unless you want to build it from source, which might provide a slight boost to the execution speed). The vignettes explain in detail how to use each of the model fitting functions in rstanarm. However, the vignettes on the CRAN website
https://cran.r-project.org/web/packages/rstanarm/index.html
do not currently show the generated images, so call browseVignettes(“rstanarm”). The help(“rstarnarm-package”) and help(“priors”) pages are also essential for understanding what rstanarm does and how it works. Briefly, there are several model-fitting functions:
- stan_lm() and stan_aov(), which just calls stan_lm(), use the same likelihood as lm() and aov() respectively but add regularizing priors on the coefficients
- stan_polr() uses the same likelihood as MASS::polr() and adds regularizing priors on the coefficients and, indirectly, on the cutpoints. The stan_polr() function can also handle binary outcomes and can do scobit likelihoods.
- stan_glm() and stan_glm.nb() use the same likelihood(s) as glm() and MASS::glm.nb() and respectively provide a few options for priors
- stan_lmer(), stan_glmer(), stan_glmer.nb() and stan_gamm4() use the same likelihoods as lme4::lmer(), lme4::glmer(), lme4::glmer.nb(), and gamm4::gamm4() respectively and basically call stan_glm() but add regularizing priors on the covariance matrices that comprise the blocks of the block-diagonal covariance matrix of the group-specific parameters. The stan_[g]lmer() functions accept all the same formulas as lme4::[g]lmer() — and indeed use lme4’s formula parser — and stan_gamm4() accepts all the same formulas as gamm::gamm4(), which can / should include smooth additive terms such as splines
If the objective is merely to obtain and interpret results and one of the model-fitting functions in rstanarm is adequate for your needs, then you should almost always use it. The Stan programs in the rstanarm package are better tested, have incorporated a lot of tricks and reparameterizations to be numerically stable, and have more options than what most Stan users would implement on their own. Also, all the model-fitting functions in rstanarm are integrated with posterior_predict(), pp_check(), and loo(), which are somewhat tedious to implement on your own. Conversely, if you want to learn how to write Stan programs, there is no substitute for practice, but the Stan programs in rstanarm are not particularly well-suited for a beginner to learn from because of all their tricks / reparameterizations / options.
Feel free to file bugs and feature requests at
https://github.com/stan-dev/rstanarm/issues
If you would like to make a pull request to add a model-fitting function to rstanarm, there is a pretty well-established path in the code for how to do that but it is spread out over a bunch of different files. It is probably easier to contribute to rstanarm, but some developers may be interested in distributing their own CRAN packages that come with precompiled Stan programs that are focused on something besides applied regression modeling in the social sciences. The Makefile and cleanup scripts in the rstanarm package show how this can be accomplished (which took weeks to figure out), but it is easiest to get started by calling rstan::rstan_package_skeleton(), which sets up the package structure and copies some stuff from the rstanarm GitHub repository.
On behalf of Jonah who wrote half the code in rstanarm and the rest of the Stan Development Team who wrote the math library and estimation algorithms used by rstanarm, we hope rstanarm is useful to you.
Also, Leon Shernoff pointed us to this post by Wayne Folta, delightfully titled “R Users Will Now Inevitably Become Bayesians,” introducing two new R packages for fitting Stan models: rstanarm and brms. Here’s Folta:
There are several reasons why everyone isn’t using Bayesian methods for regression modeling. One reason is that Bayesian modeling requires more thought . . . A second reason is that MCMC sampling . . . can be slow compared to closed-form or MLE procedures. A third reason is that existing Bayesian solutions have either been highly-specialized (and thus inflexible), or have required knowing how to use a generalized tool like BUGS, JAGS, or Stan. This third reason has recently been shattered in the R world by not one but two packages:
brmsandrstanarm. Interestingly, both of these packages are elegant front ends to Stan, viarstanandshinystan. . . . You can install both packages from CRAN . . .
He illustrates with an example:
mm <- stan_glm (mpg ~ ., data=mtcars, prior=normal (0, 8))
mm #===> Results
stan_glm(formula = mpg ~ ., data = mtcars, prior = normal(0,
8))
Estimates:
Median MAD_SD
(Intercept) 11.7 19.1
cyl -0.1 1.1
disp 0.0 0.0
hp 0.0 0.0
drat 0.8 1.7
wt -3.7 2.0
qsec 0.8 0.8
vs 0.3 2.1
am 2.5 2.2
gear 0.7 1.5
carb -0.2 0.9
sigma 2.7 0.4
Sample avg. posterior predictive
distribution of y (X = xbar):
Median MAD_SD
mean_PPD 20.1 0.7
Note the more sparse output, which Gelman promotes. You can get more detail with
summary (br), and you can also useshinystanto look at most everything that a Bayesian regression can give you. We can look at the values and CIs of the coefficients withplot (mm), and we can compare posterior sample distributions with the actual distribution with:pp_check (mm, "dist", nreps=30):
This is all great. I’m looking forward to never having to use lm, glm, etc. again. I like being able to put in priors (or, if desired, no priors) as a matter of course, to switch between mle/penalized mle and full Bayes at will, to get simulation-based uncertainty intervals for any quantities of interest, and to be able to build out my models as needed.
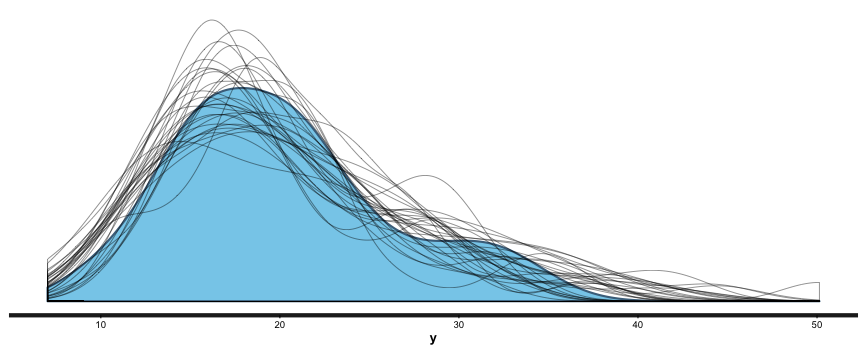
This is awesome.
Awesome, indeed! Especially that it’s already being used (we got some hint of that from people using early development versions). It shows that Andrew and Ben were right about the demand.
“ARM” is for “applied regression modeling.”
After the usual bike shed debate, we settled on “RStan” in writing and “
rstan” in R code, “Stan” in writing and “stan” in C++ code, so I suppose this will be “RStanARM” in writing and “rstanarm” in R code. Unless you’re Andrew. Andrew pronounces the acronym “ARM” like the English noun or verb “arm”, so if he sticks to his conventions, he’ll write/type/LaTeX it as a proper name (i.e., “Arm”).That sounds like a bad idea. When you mean the R package, I think you should write its name faithfully no matter if you are writing it in R code or elsewhere, otherwise your readers may try to install.packages(“RStanARM”) or library(RStanARM). Worse, they may never remember if the package name was rstanarm, RstanArm, RStanArm, RSTANARM, …
This is pretty standard for software. For instance, you guys write “R Markdown” and “Shiny” in text rather than “rmarkdown” and “shiny” when talking about the systems. In your defense, I haven’t seen that for knitr (except on Wikipedia), so I try not to start a sentence with “knitr”!
Java is written “Java” but in code the top-level namespace is “java”. Apache Lucene in code is “apache.lucene”.
Yes, when we talk about the system, we write R Markdown and Shiny. But I was saying “when you mean the package”. In this blog post, I think rstanarm mostly means the R package. I never say “the Shiny package, or the R Markdown package”, and I think it is a bad idea to say “the RStanARM package”. I’m very careful with the name knitr. You are very sharp to have observed that :) I don’t have a single instance of “Knitr” in my book or elsewhere except in Wikipedia (I don’t have a choice there). On the other hand, Stan sounds more like a system than rstanarm, so I’m okay with Stan but not so with RStanARM (to be clear, I’m not judging the value of the latter).
Okay okay, enough bikeshedding for me now. Please write in whatever fashion you like. I was here just to show the bike-shed effect :-D
This is very awesome. Also includes Shiny functions, so you can interactively scrutinize your models! :-D
Looks great.
We have been using this package with a book publisher and it helped with accelerating the model development process. We usually start with a model in rstanarm, build a few of them quickly, and then build up a model in Stan when we need to include things that the package does not yet support, like MA process for errors, etc.
Played with this well unto late at night. Really great for “users” like me. I have the direct comparison with trying to write a multilevel model by hand, with glmer2stan and now this.
Before this release, I sometimes dared to use Stan etc., now I hope to make it my default go-to.
Now I’m hoping for someone doing a nice automated function for marginal effect plots and a bit more extractors for people who prefer other to customise their plotting/do it somewhere else.
It is not fully automated, but you can convey marginal effects by plotting the posterior_predict() output with the newdata argument being a data.frame that sets all other predictors to whatever level and letting the variable of interest vary over the relevant range.
Thanks Ruben, glad you like it! Like Ben said, it’s not too tough to make the kind of plots that (I think) you’re referring to after calling posterior_predict, but we do have some more plotting functions along these lines that we’re working on for future releases. Also, what type of “extractors” are you referring to?
Yes, I tried to roll my own marginal effect plots for one model yesterday. It seemed fairly easy, but hard enough for me to make a mistake. I didn’t get the intervals right, maybe it was too late for me. And too early now ;-) I used stan_lmer and re.form = ~0. The predicted lines were wiggly (not straight) and the intervals were wider than what I expected. I wanted credible intervals for the marginal effects in my models, but I think I got prediction intervals for out-of-sample prediction. In predict.MCMCglmm you could toggle this with the interval argument.
Functions like plot(allEffects(fit)) or sjPlot::sjp.int(fit, type = ‘eff’) make this super convenient, because they do the newdata generation for me and automatically pick appropriate levels for the moderator. That makes them one of my go-tos for exploring models myself and explaining models to others.
With regard to extractors: the default plotting functions are great (and even greater after I realised you used ggplot2 and just made it look more like base plots, so I could easy increase the font size after generation), they generate some underlying data that could be even easier to get.
For example I often want to compare coefficients from different models and put them in a new coefficient plot (eg. for robustness checks).
For now, I can do
coefs = plot(fit, “stan_plot”, pars = “beta”)
rbind(coefs$data, coefs2$data)
and then pick names that make it comparable to other packages’ terminology. But maybe unifying that sort of thing is more something for broom::tidy.
On a different note your plots don’t respect the ggplot2 theme I set. I guess you have more solid ideas on how to plot right, but since I know the environment I will be plotting in (e.g. a knitr document with a certain width/height set), I would like the ability to set defaults.
But don’t let that distract you, I’m mostly amazed.
Just now I noticed that you give me progress reports when running chains on multiple cores. That those are usually absent is my main (mostly psychological) reason to prefer single cores sometimes.
Thanks for the detailed response. This kind of feedback is great. I’ll have to think about all this a bit and see what it makes sense to do for rstanarm. We should definitely submit a pull request to the broom package with “tidy” functionality for rstanarm models. That should be pretty easy to do.
Great! This is a step towards statistical heaven. It opens wider the doors to the dungeon and the loft of statistics. In the dungeon, you can easily use Bayesian credible intervals whenever you fail to get confidence intervals. In the loft, you can do asymptotically exact hypothesis tests and predictions of likely future outcomes for the same model with limited efforts. To access statistical heaven, it remains to construct methods to easily do hypothesis tests that remain exact when you have only a limited amount of data and must use non-trivial models.