Someone pointed me to this post by “Neuroskeptic”:
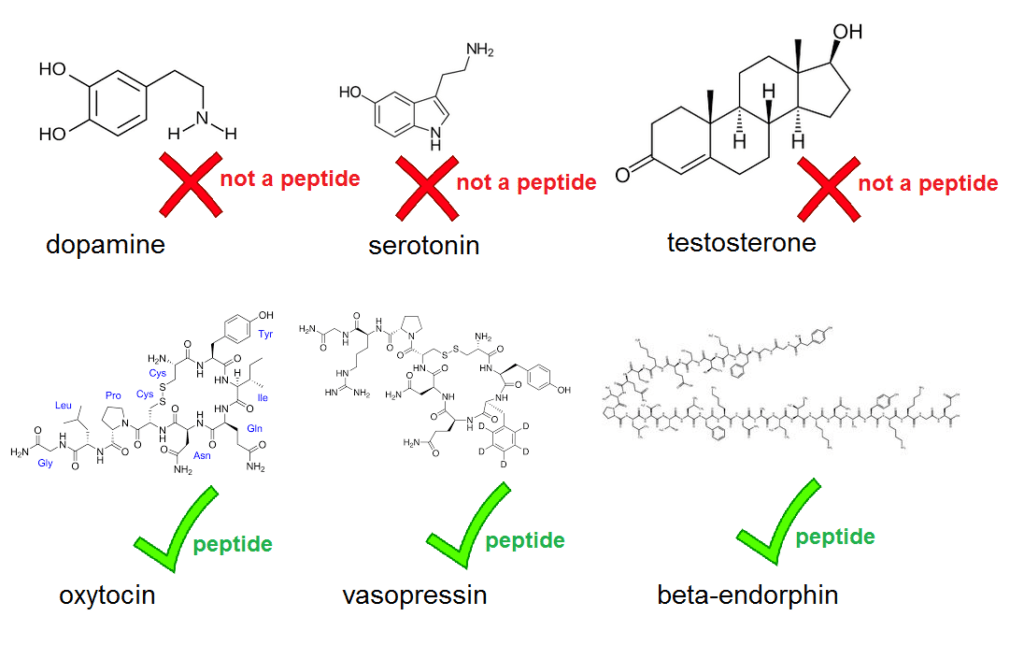
A new paper in the prestigious journal PNAS contains a rather glaring blooper. . . . right there in the abstract, which states that “three neuropeptides (β-endorphin, oxytocin, and dopamine) play particularly important roles” in human sociality. But dopamine is not a neuropeptide. Neither are serotonin or testosterone, but throughout the paper, Pearce et al. refer to dopamine, serotonin and testosterone as ‘neuropeptides’. That’s just wrong. A neuropeptide is a peptide active in the brain, and a peptide in turn is the term for a molecule composed of a short chain of amino acids. Neuropeptides include oxytocin, vasopressin, and endorphins – which do feature in the paper. But dopamine and serotonin aren’t peptides, they’re monoamines, and testosterone isn’t either, it’s a steroid. This isn’t a matter of opinion, it’s basic chemistry.
The error isn’t just an isolated typo: ‘neuropeptide’ occurs 27 times in the paper, while the correct terms for the non-peptides are never used.
Neuroskeptic speculates on how this error got in:
It’s a simple mistake; presumably whoever wrote the paper saw oxytocin and vasopressin referred to as “neuropeptides” and thought that the term was a generic one meaning “signalling molecule.” That kind of mistake could happen to anyone, so we shouldn’t be too harsh on the authors . . .
The authors of the papers work in a psychology department so I guess they’re rusty on their organic chemistry.
Fair enough; I haven’t completed a chemistry class since 11th grade, and I didn’t know what a peptide is, either. Then again, I’m not writing articles on peptides for the National Academy of Sciences.
But how did this get through the review process? Let’s take a look at the published article:

Ahhhh, now I understand. The editor is Susan Fiske, notorious as the person who opened the gates of PPNAS for the articles on himmicanes, air rage, and ages ending in 9. I wonder who were the reviewers of this new paper. Nobody who knows what a peptide is, I guess. Or maybe they just read it very quickly, flipped through to the graphs and the conclusions, and didn’t read a lot of the words.
Did you catch that? Neuroskeptic refers to “the prestigious journal PNAS.” That’s PPNAS for short. This is fine, I guess. Maybe the science is ok. Based on a quick scan of the paper, I don’t think we should take a lot of the specific claims seriously, as they seem to based on the difference between “significant” and “non-significant.”
In particular, I’m not quite sure what is their support for the statement from the abstract that “each neuropeptide is quite specific in its domain of influence.” They’re rejecting various null hypotheses but I don’t know that this is supporting their substantive claims in the way that they’re saying.
I might be missing something here—I might be missing a lot—but in any case there seem to be some quality control problems at PPNAS. This should be no surprise: PPNAS is a huge journal, publishing over 3000 papers each year.
On their website they say, “PNAS publishes only the highest quality scientific research,” but this statement is simply false. I can’t really comment on this particular paper—it doesn’t seem like “the highest quality scientific research” to me, but, again, maybe I’m missing something big here. But I can assure you that the papers on himmicanes, air rage, and ages ending in 9 are not “the highest quality scientific research.” They’re not high quality research at all! What they are, is low-quality research that happens to be high-quality clickbait.
OK, let’s be fair. This is not a problem unique to PPNAS. The Lancet publishes crap papers, Psychological Science published crap papers, even JASA and APSR have their share of duds. Statistical Science, to its eternal shame, published that Bible Code paper in 1994. That’s fine, it’s how the system operates. Editors are only human.
But, really, do we have to make statements that we know are false? Platitudes are fine but let’s avoid intentional untruths.
So, instead of “PNAS publishes only the highest quality scientific research,” how about this: “PNAS aims to publish only the highest quality scientific research.” That’s fair, no?
P.S. Here’s a fun little graphics project: Redo Figure 1 as a lineplot. You’ll be able to show a lot more comparisons much more directly using lines rather than bars. The current grid of barplots is not the worst thing in the world—it’s much better than a table—but it could be much improved.
P.P.S. Just to be clear: (a) I don’t know anything about peptides so I’m offering no independent judgment of the paper in question; (b) whatever the quality is of this particular paper, does not affect my larger point that PPNAS publishes some really bad papers and so they should change their slogan to something more accurate.
P.P.P.S. The relevant Pubpeer page pointed to the following correction note that was posted on the PPNAS site after I wrote the above post but before it was posted:
The authors wish to note, “We used the term ‘neuropeptide’ in referring to the set of diverse neurochemicals that we examined in this study, some of which are not peptides; dopamine and serotonin are neurotransmitters and should be listed as such, and testosterone should be listed as a steroid. Our usage arose from our primary focus on the neuropeptides endorphin and oxytocin. Notwithstanding the biochemical differences between these neurochemicals, we note that these terminological issues have no implications for the significance of the findings reported in this paper.”
Candidate gene studies in small (i.e., <10,000) studies still being published? In PNAS? Without preregistration? Hard to believe.
This reminds me of the Tower of Babel. Neuropeptide instead of neurotransmitter, accuracy instead of AUC[1], another recent one is microcephaly instead of micrencephaly[2], etc. Keep your eye on the accuracy confused with AUC issue as ML/AI becomes more common. Area under the ROC curve seems like fertile territory for confusion:
https://en.wikipedia.org/wiki/Receiver_operating_characteristic#Area_under_the_curve
[1] https://techcrunch.com/2017/05/11/apples-watch-can-detect-an-abnormal-heart-rhythm-with-97-accuracy-ucsf-study-says/
[2] “Because the size of the brain is mostly determined by the size of the head, microencephaly is implied when discussing microcephaly.” – https://en.wikipedia.org/wiki/Microcephaly#Microencephaly (This is in general false. It is only true if you define microcephaly to be a head <3sd from the mean, not the usual 2sd.)
Seeing that the authors went down the typical “safe” statistical inference route (judging Figure 1), perhaps they used bar charts instead of line charts to strictly emphasise the comparison between males and females and avoid drawing unintended connections between the gene polymorphisms?
It seems likely to me that the data generating process here was:
1) First start writing about actual neuropeptides: oxytocin, vasopressin, etc…
2) after a while, start thinking about what other neuro-active molecules might have similar issues…
3) Insert information about these other molecules without realizing that it means you should go through the paper and change “neuropeptide” to “neuroactive molecule” because it’s not primarily the chemical structure that you were interested in.
4) Send paper for review to people who aren’t chemists
5) ???
6) Profit!
This doesn’t bother me as much as it would if the mislabeling led to substantive predictions that were based on the misclassification. For example, suppose I predict that neuropeptide breakdown leads to certain nitrogen compounds in the brain because of the nitrogen in peptides, and then I say that this is going to also happen for testosterone (a steroid which has no nitrogen in it). That’d be a serious substantive scientific error, whereas it seems that maybe this error in this paper could be fixed by the global search and replace of “neuropeptide” with “neuroactive molecule” and then the whole thing might be perfectly correct. Such “global search and replace” fixable mislabelings are relatively innocuous, embarassing, but not really substantive.
Maybe I’m being obtuse, but why do you always refer to this journal as PPNAS? Is this some variety of in-joke?
PPNAS is short form of prestigious journal PNAS
Most appropriate given the origin of the word prestigious ;-)
http://www.dictionary.com/browse/prestigious
1540-50; < Latin praestigiōsus full of tricks, deceitful, equivalent to praestigi(um) (see prestige ) + -ōsus -ous
(Word origin originally from Susan Haack https://en.wikipedia.org/wiki/Susan_Haack)
Thanks for that one Keith, I had no idea this word was so loaded. I found this document to contain an interesting historical account. According to them, prestige was all about playing shell games, dulling the intellect, “effectiveness not comprehensible by logic”, domination of an idea that “paralyzes our critical faculty”, “authority which stops all discussion or with a protecting pathos which precludes criticism”, etc.
I think we found what the “p” in “significant p-value”* stands for!
* Usual disclaimer: p-values are probably fine when used to test a real model/hypothesis rather than a default strawman one
An equally accurate and revealing name would be PCNAS
Michael:
As I wrote in my post, I’m sure that PPNAS publishes lots of great articles. My problem is not with the fact that it publishes the occasionally obviously bad paper—this is essentially unavoidable given that (a) the journal publishes 3000 papers per year, and (b) Susan Fiske has the authority to accept articles. My problem is with their statement that they publish “only the highest quality scientific research,” a statement which everyone other than Fiske must know is untrue.
I hate it when people say things that they know are false. Sure, there’s a place in life for white lies, but not so much in science!
I understand, and given your important perspective and site, “PPNAS” is a better jab. But knowing what gets published there and what doesn’t even get sent out for review, there is a certain point of view, even more evident than in mainstream psychology journals. For example, a friend got a paper sent back without review on police shootings and race (the paper found no evidence of racial bias) with the explanation that “The topic does not have broad appeal.” (Right, no one is interested in racial bias in police shootings.) Then recently this: http://www.pnas.org/content/114/25/6521.full . (Some interesting questions regarding potential forking paths here.)
PCNAS
Michael:
I have the impression that Lancet is politically biased too, as for example in that paper from 2006 on the Iraq survey, or that paper from last year on gun control.
From the other direction, econ journals can be biased too. For example we submitted this paper to an econ journal which rejected it, partly on the grounds that the conclusion didn’t fit their model of the world.
It’s a tough call—to some extent, if a statistically-obtained empirical result disagrees with your theory, you should be questioning. But at some point it does seem to edge into ideological bias.
In the Fiske-accepted papers (air rage, himmicanes, ages ending in 9, etc.), I wouldn’t say there’s any political angle, but there is, one could say, a scientific ideology in place, which is that small, seemingly meaningless inputs can have large, predictable effects on behavior. I’ve criticized this view on the blog for awhile and will continue to do so.
> Sure, there’s a place in life for white lies, but not so much in science!
And there is a place for science which seems less and less likely to be in journals…
OK, occasionally it can sneak in ;-)
It’s also a little suspicious that in the very first figure, first plot the SNP rs61525 doesn’t exist in the Supplemental data where it’s supposed to be. I would hazard a guess they meant rs6152.
Pen pineapple apple science?
Andrew wrote: ” I wonder who were the reviewers of this new paper.”
Could it be that Fiske reviewed it herself and didn’t send it out to anyone else for review? (The reason this seems plausible to me is that once, many years ago, I submitted a paper to the Proceedings of the American Mathematical Society and got an extremely quick reply saying it had been accepted, so suspected that the editor to whom I submitted it reviewed it himself. If I remember correctly, it was a paper that had a serious error in it, which someone I had sent a preprint to pointed out to me, at which point I withdrew the paper before it was published.)
> which someone I had sent a preprint to pointed out to me
Good call – as soon as you suspect an accepted paper was not adequately reviewed you know you are at risk.
Neuroskeptic’s comments apparently reached the authors because the PNAS website has a correction on the paper from the authors, the gist of which is that they were indeed wrong in their application of the term “neuropeptide,” but it doesn’t materially alter their conclusions.
Several other comments:
1. I noticed the same thing about the SNP labeled as rs61525 in Figure 1 that awinter mentioned above. It should be rs6152 (they labeled it correctly in the SI appendix).
2. There are two other things that are somewhat odd about the way that the effects in the first row of graphs in Figure 1 are represented:
a) The functional molecule is labeled as Testosterone. rs6152 is a SNP in the AR (Androgen Receptor) gene (note: by convention gene names are supposed to be italicized to distinguish them from their protein products. I wasn’t sure if I could include html in this comment, so my references to the gene, AR, are not italicized). While it is true that the androgen receptor binds testosterone, it also binds other endogenous androgens. The assumption that a variant in the AR gene effectively acts as a proxy for testosterone levels might have some validity, but it would have been preferable just to label that first row as AR effects in the absence of direct measurement of testosterone levels in the subjects.
b) The x-axes in this first row of Figure 1 are repeatedly labeled “Acarrier” and “GG.” This is because rs6152 involves a mutation of the ancestral G allele at that base position to an A allele leaving two different alleles in this population. The authors are effectively testing whether carrying any A alleles (genotypes AA and AG) produces a phenotypic difference from carrying no A alleles (genotype GG). The AR gene is on the X chromosome. The authors state in the Appendix SI that rs6152 is a “haploid” SNP. It’s the gene that is haploid (a single copy of the gene on one chromosome) as opposed to diploid (the normal complement of two copies of the gene, one copy on each of the two homologous chromosomes). The gene is only haploid in males because they have only one X chromosome. Females, because they typically have two X chromosomes, are diploid for this gene. So, indeed, the females in the study could have genotypes AA, AG or GG. Males can only have genotypes A or G based on the rs6152 variant. Yet, in Figure 1 that contains separate bars for males and females, males are shown as being either Acarriers (possible) or having the genotype GG (impossible). It is true that in any given cell in females, one of the two X chromosomes is silenced (X-inactivation), but which of the two X chromosomes (maternal or paternal) is inactivated appears to be random. Nonetheless, this does not render females haploid for genes on the X chromosome. Referring to the AR gene gene as being haploid is technically only true for males.
3. Methodologically, the authors make some confusing statements in the Analysis section of Appendix SI. They used PLINK to analyze their data. In particular, they said that they a) pruned the set of SNPs for each gene based on Linkage Disequilibria among the SNPs, and b) they used a permutation procedure (N = 1000 permutations per genotype/phenotype association test) to control Type I error inflation from multiple testing rather than using a Bonferroni correction.
Regarding (a): Linkage Disequilibrium refers to the non-independence between alleles at locations close to each other on the same physical chromosome. Without going into the mechanisms that produce or disrupt LD, suffice it to say that SNPs that are close to each other tend to have high LD. In statistical terms this means that knowing the genotype at one location allows you to predict with better than chance accuracy, perfect accuracy in case of perfect LD, the genotype at a nearby location. So two SNPs that are close to each other will not act as independent predictors of the phenotype (nor will they be passed on independently in the germline). Pruning based on high LD means removing SNPs from the analysis so that one is left essentially with a set of SNPs that are in Linkage Equilibrium; these SNPs (actually, the corresponding genotypes) then act as approximately independent predictors of the phenotype. The implications of this pruning are discussed after addressing PLINK’s documentation on permutation testing.
Regarding (b): PLINK’s documentation (v 1.07) says that using permutation testing to control Type I error given multiple tests of significance is less stringent than a Bonferroni correction “Because the permutation schemes preserve the correlational structure between SNPs … in comparison to the Bonferroni, which assumes all tests are independent.”
Now Pearce et al. say “Although genotypic model significance levels did not always survive correction for multiple tests using mperm, the consensus is that these corrections are likely to be overly conservative because they fail to account for dependence between tests due to linkage disequilibrium…” This appears to directly contradict what PLINK’s documentation says in two ways: 1) PLINK says that this procedure is less stringent than Bonferroni, not an overly conservative correction as Pearce et al. said. 2) PLINK’s documentation notes that the permutation schemes “preserve the correlational structure between SNPs.” Yet, Pearce et al. claim that these permutation schemes fail to take Linkage Disequilibrium into account, directly contradicting the PLINK documentation (and from the description of how the permutation schemes are implemented, it is clear that when multiple SNPs are tested simultaneously for association with a phenotype, the SNPs are not permuted; it is basically only the phenotypes that get randomly re-assigned to subjects, who, in turn, maintain all of their genotypes on the various SNPs).
The “kicker” in all of this, however, is that Pearce et al. already pruned the set of SNPs within each gene down to a set that was in approximate Linkage Equilibrium; that is, a set of nearly independent genotypic predictors. Retaining associations that failed to survive the multiple testing corrections because the corrections were overly conservative in not taking into account the correlational structure among the SNPs makes no sense if there is no non-zero correlational structure in the first place.
Finally, the pruning procedure that they used left many of the candidate genes being represented only by one SNP in the analysis. Of course, there can be no correlational structure among the genotypic predictors if there is only one predictor.
“1) PLINK says that this procedure is less stringent than Bonferroni, not an overly conservative correction as Pearce et al. said”
It’s very easy to be both less stringent than Bonferroni, yet still overly conservative. Bonferroni preserves family-wide type 1 error rates in the worst possible case. The assumption of independence allows for much powerful corrections that preserve type 1 error rates…but are still overly conservative if the tests are actually positively correlated.
Thanks. I failed to appreciate that distinction because I was focused on the inconsistencies in their argument for retaining effects that failed to survive the correction for multiple testing. I should have been clearer that I didn’t buy their argument because it was a) inconsistent with what they had done (retained SNPs for each gene that were explicitly independent or nearly so), b) contradicted by the way that PLINK describes the permutation schemes, and c) moot for many genes because they only used a single marker per gene.